Our Research
Our group focuses on generating hierarchical functional soft materials using synthetic polymers, peptides, proteins, small organic molecules and nanoparticles as building blocks. Our projects are split into 3 subgroups.
Overview
Future technologies depend on the development of functional materials having: (1) hierarchical structures spanning multi-length scale down to the molecular level, (2) built-in functionalities (biological, optical, electronic, magnetic properties), (3) superior selectivity with sensitivities comparable to what found in nature, (4) responsiveness to external stimuli. These require one to select the right building blocks, understand the principles underpinning the self-assembly, and use these principles to direct the assemblies at various length scales to obtain targeted functional materials. More importantly, we need to develop a versatile methodology to generate new functional materials simply by substituting building blocks, instead of re-building them from scratch. Numerous building blocks have been explored to achieve this end and obtaining functional materials to meet these requirements remains a significant challenge to the soft material community.
Our group focuses on generating hierarchical functional soft materials using synthetic polymers, peptides, proteins, small organic molecules and nanoparticles as building blocks. Each offers unique properties and are complimentary to each other. Natural proteins have complexities in terms of structure and functionaly unmatched by synthetic materials. De novo designed peptides are minimalistic natural proteins that mimic natural protein functionalities but are subject to degradation. Small organic molecules, especially those with optical, electronic and magnetic properties, can be readily synthesized with molecular control to provide built-in functionalities. Nanoparticles, due to their size, exhibit unique properties not seen in macroscopic materials and constitute essential building blocks in the fabrication of nanodevices. It is non-trivial to obtain macroscopic assemblies of either small molecule or nanoparticle at low cost. Polymers with different architectures can be synthesized and are amenable to various processing techniques. Developments in polymer sciences provide guidance to manipulate their assemblies at different length scales. However, generating molecular level assemblies with designer functionalities using polymers alone is challenging. Synergistic assemblies of these selected building blocks clearly have tremendous potential to construct technologically important functional materials. By developing fundamental understanding of the physics of assemblage, our group aims to generate hierarchical structures spanning multi-length scales down to few nanometers with built-in biological, electrical and magnetic functionalities.
Currently, our group is engaged in three major research activities to address the challenge of multi-length scale, hierarchical functional assemblies using these building blocks, split into 3 distinct subgroups.
Nanocomposites
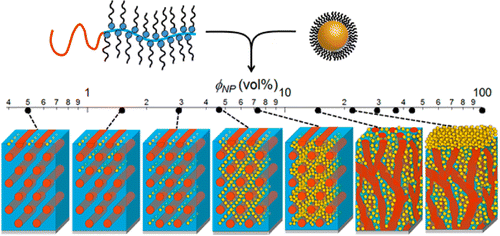
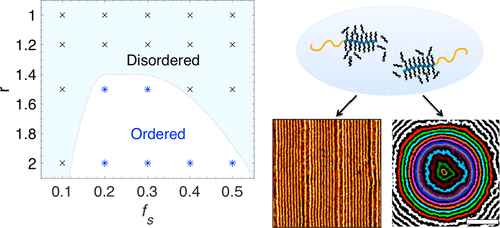
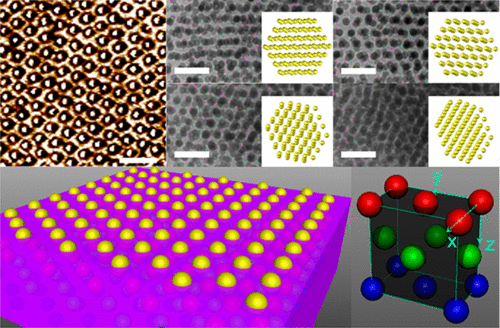
Self-assembling nanocomposites form ordered structures on nanometer length-scales, overcoming the limits of traditional manufacturing processes and introducing new collective properties. The Nanocomposite subgroup specializes in precise and tunable self-arrangement of inorganic nanoparticles within soft matrix materials. Much of our work has focused on nanoparticle assembly using a supramolecule of polystyrene-b-poly(4-vinyl pyridine) with small molecule 3-pentadecylphenol, abbreviated PS-P4VP(PDP). The small molecule has a large impact on assembly kinetics and thermodynamic equilibria, and this system can achieve morphologies that are inaccessible to a simple diblock copolymer matrix.
The PS-P4VP(PDP) supramolecule system has been studied in bulk and in thin films, with many different nanoparticle shapes, compositions, and arrangements. Other current work in the Nanocomposite subgroup includes controlled assembly of polymer-grafted nanoparticles into superlattices, 3D-printing of nanocomposites, and conductive nanocomposite spray-coatings. Ultimately, we hope to decode the design rules for a rich library of nanocomposites, enabling applications in sensing, optics, electronics, and energy storage.
Protein-based functional materials
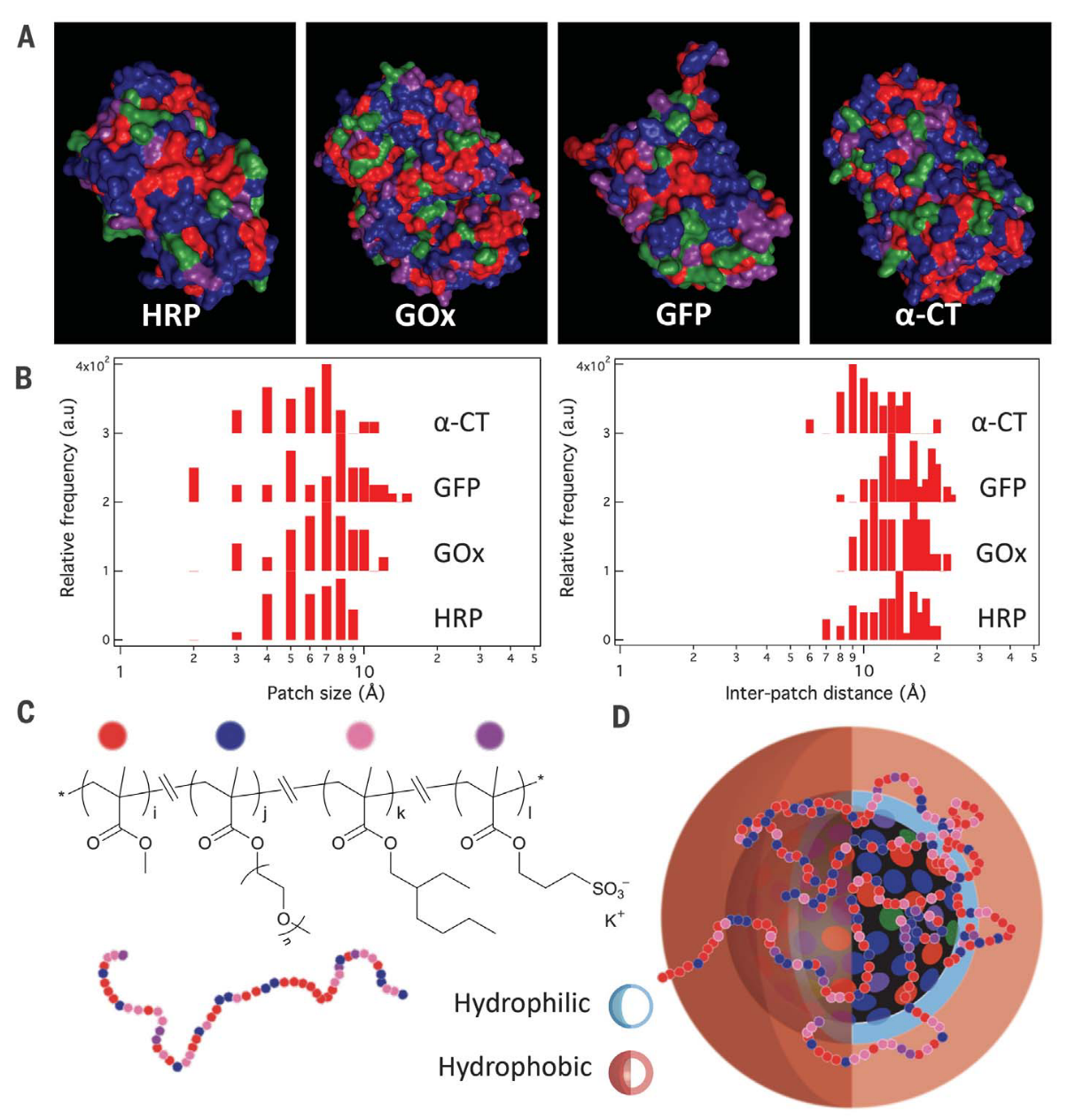
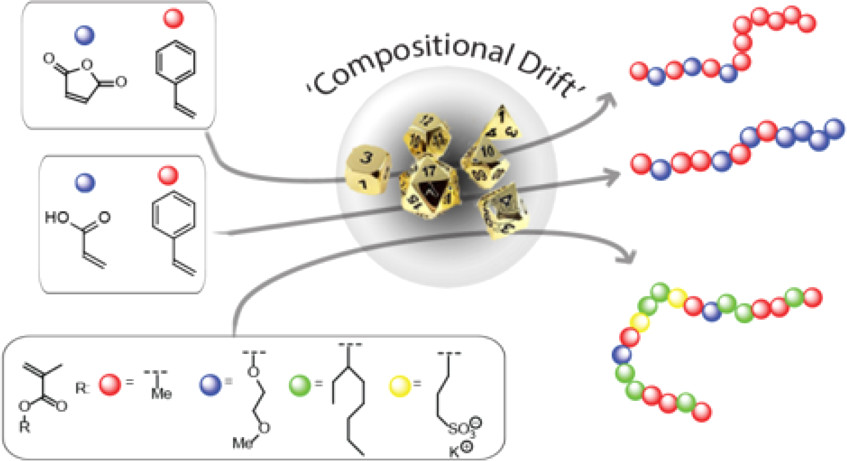
In nature, intrinsically disordered proteins fold into local chain conformations, yielding surfaces that are chemically diverse and heterogeneous. Our research focuses on designing amphiphilic, statistically random heteropolymers (RHP) that mimic these disordered proteins to offer versatile approaches for various synthetic and hybrid-biomaterial applications. We designed a four-monomer RHP that can act as a chaperone-like polymeric shell to surround, solubilize, and stabilize proteins in a wide variety of non-native environments. This RHP approach allows us to incorporate active proteins into synthetic polymers to create a new class of materials with functions found only in living systems. By optimizing composition and statistical monomer distributions we can enable the cell-free synthesis of membrane proteins with proper folding for transport and enzyme-containing plastics for toxin bioremediation.
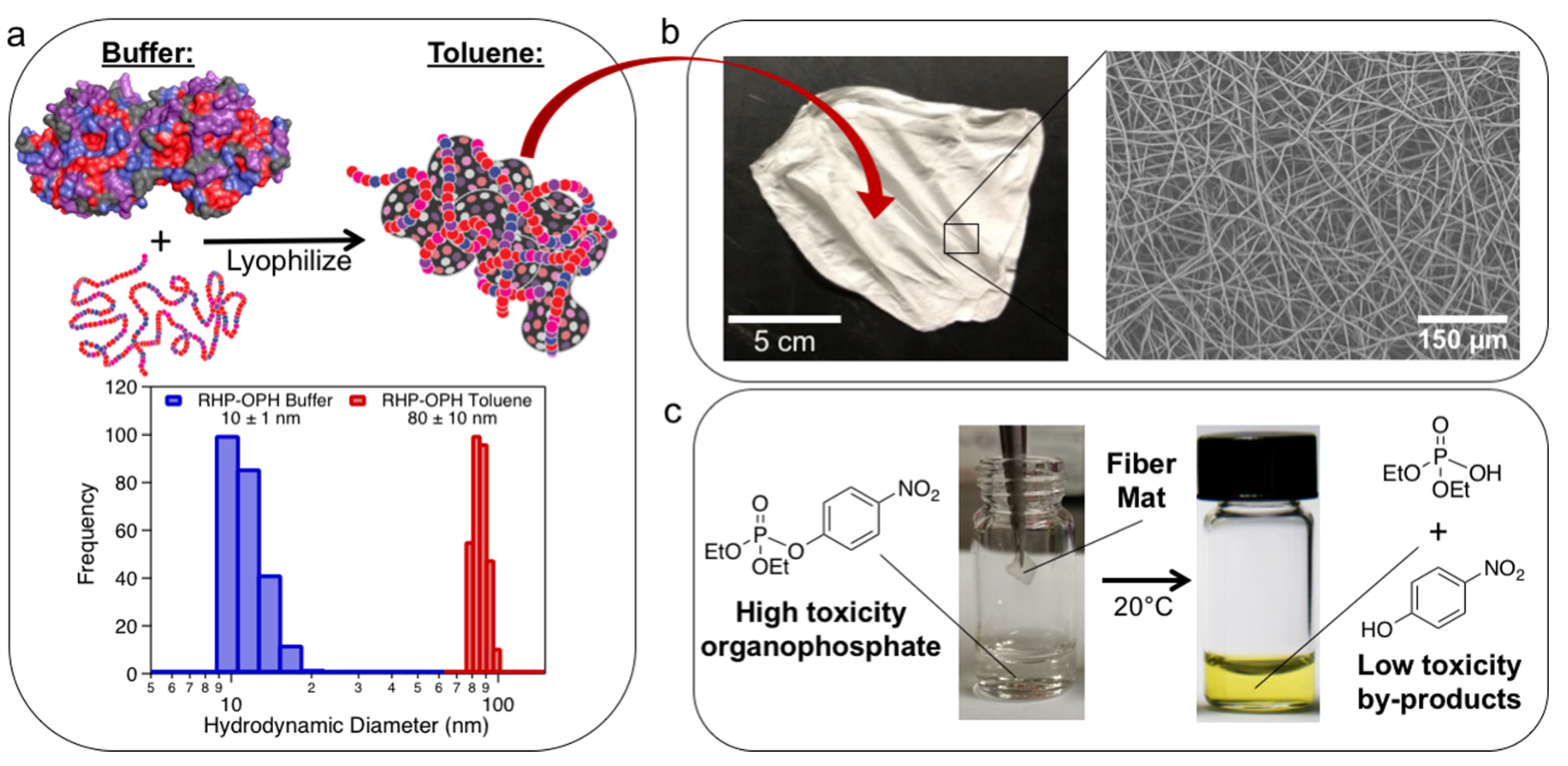
A specific example application we have shown is for toxin bioremediation of organophosphates, acutely toxic chemicals found in pesticides and chemical warfare agents. Solving the problem of inherent enzyme fragility in non-native environments while retaining high catalytic efficiency, selectivity and sensitivity allowed us to fabricate highly effective, reusable, non-toxic and portable organophosphorus hydrolase loaded fiber mats capable of degrading toxic organophosphates over a broad range of relevant concentrations (from 8 to 8250 ppm)
Our projects span across the entire soft materials development pipeline from developing novel software for guided polymer design to RAFT synthesis optimization to post-functionalization to applications inclusive of neurotoxin remediation, metal binding and gas barrier materials.
Rational Design of Nanocarriers for Tumor Therapy
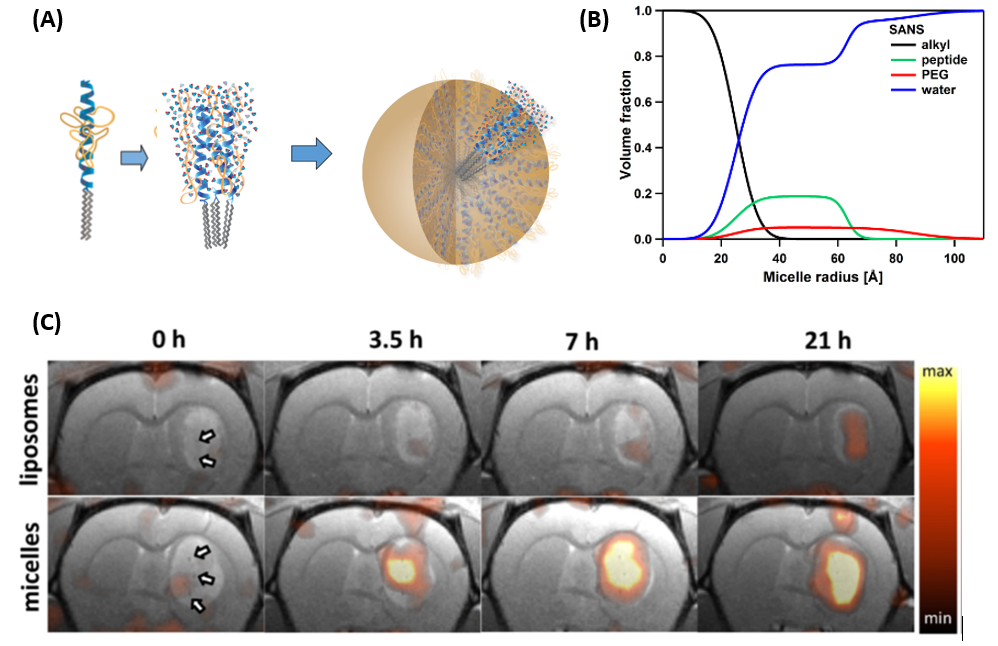
Our lab has developed a delivery platform that exploits polymer entropic repulsion directed by the protein tertiary structure of the head group of an amphiphile to generate long-circulating, ~15-20 nm self-assembled nanoparticles with desirable biodistribution and tunable stability. Recent work investigating the internal structure of the 3-helix micelle (3HM) system has kindled interest in the role of the structural properties of these particles in the enablement and mechanism of enhanced in vivo transport to, and within, the tumor microenvironment.
Based on small-angle neutron scattering (SANS) studies, 3HM contains ~85% v/v water distributed outside of the alkyl tail core of the particle, residing mainly in the peptide shell layer and polyethylene glycol (PEG) corona. The high hydration level coupled with self-assembled structure is theorized to impart flexibility, which can be beneficial for extravasation through narrow gap junctions and penetration through tortuous extracellular matrix. Planar hydrophobic drugs such as Doxorubicin can be intercalated among alkyl groups in the core, making 3HM an attractive nanocarrier for tumor therapy, and ongoing studies reveal extended internalization into patient-derived glioblastoma (GBM) cells. Considering that systemically injected 3HM accumulates significantly more than liposomes within orthotopic GBM xenograft tumor, 3HM-DOX appears to be a viable therapy for poorly-permeable tumors such as GBM.